Synthetic lethality in cancer
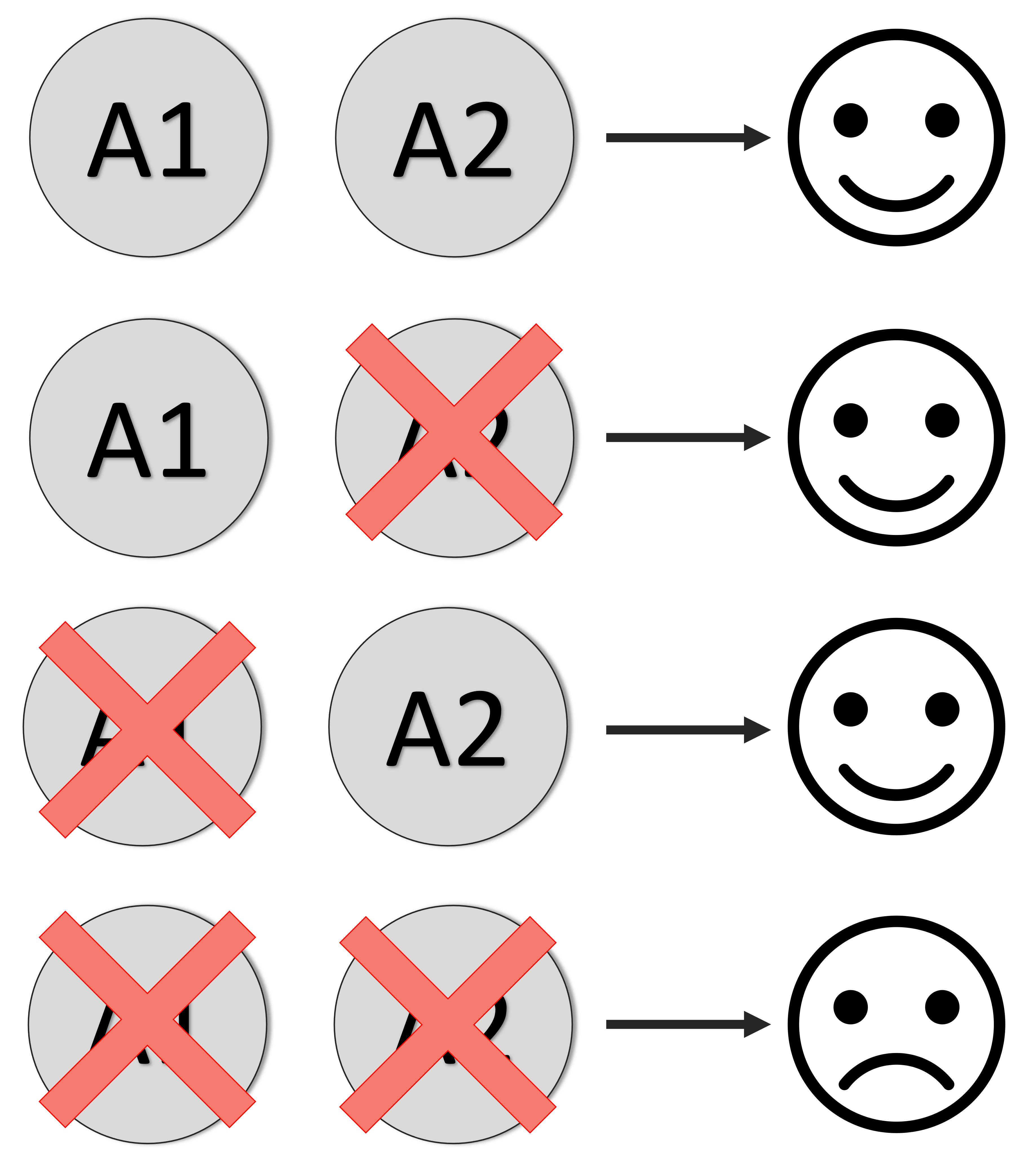
Synthetic lethal interactions are identified when two genes can be perturbed individually with little or no fitness consequence but their combined perturbation results in cell death. In the context of cancer, synthetic lethal interactions can be exploited for the development of targeted therapeutics – targeting a synthetic lethal partner of a tumour suppressor gene can be a means to selectively kill tumour cells without harming healthy cells. The first therapies based on synthetic lethal interactions are now used in the clinic and large-scale experimental efforts are now underway to identify new synthetic lethal interactions. A major challenge to the identification of new synthetic lethal interactions is that they appear to be highly context specific – an interaction between two genes may be synthetic lethal only in a specific cell line or specific cancer type. We have previously developed computational approaches to identify robust synthetic lethal interactions – those that operate across multiple contexts or cell lines. We are now interested in understanding what makes a gene pair likely to be synthetic lethal in a specific context and in developing machine learning models to predict in what contexts a given gene pair are likely to be synthetic lethal.
The contribution of paralogs to the genetic robustness of tumours
Tumour cells tolerate enormous numbers of genetic perturbations, including loss-of-function mutations and homozygous deletions of entire protein-coding genes. Despite these perturbations, tumour cells continue to function and indeed thrive. This ability of cells and organisms to tolerate genetic perturbations is generally called genetic robustness. In model organisms, paralogs (duplicate genes) have been shown to be crucial to genetic robustness. Because they are derived from the same ancestral gene, paralogs often share significant sequence similarity and in some cases functional similarity. Pairs of paralogs can therefore often compensate for each others’ loss, allowing one member of the paralog pair to be mutated or deleted with relatively little fitness consequence. We are interested in understanding how these compensatory relationships between paralogs shape tumour genome evolution, how they impact the vulnerabilities of cancer cell lines, and how they facilitate the rewiring of molecular interaction networks in cancer. There is a connection here to our work on synthetic lethality – compensatory relationships between paralogs are often revealed by synthetic lethal interactions and so we have developed an approach to systematically predict synthetic lethality between paralogs.
Network rewiring in response to genetic perturbations in cancer
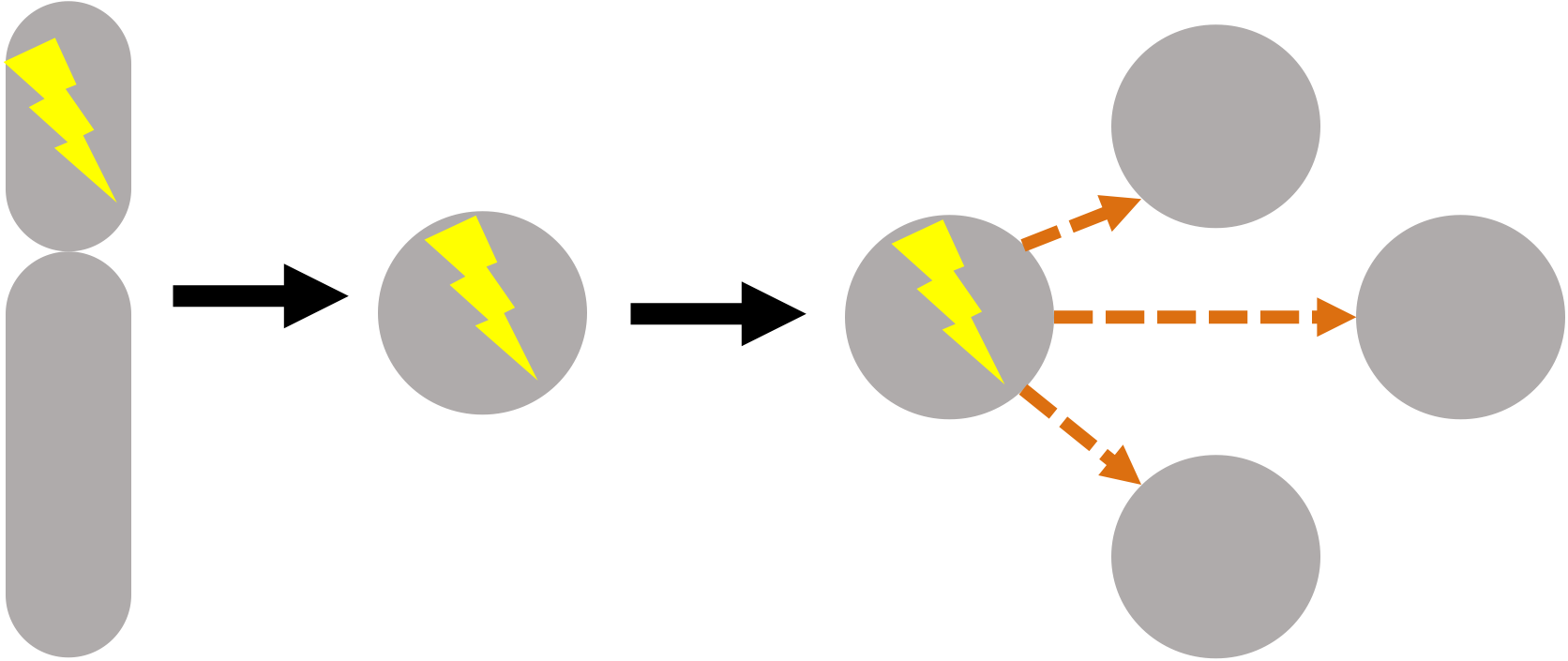
Genes, and their protein products, work in dense interconnected networks. A consequence of this is that the mutation of one gene in cancer often propagates through molecular interaction networks. For instance, the mutation of one member of a protein complex can result in altered protein abundance of the entire protein complex. Likewise, the mutation of a transcription factor can result in the altered expression of many of that transcription factor’s targets. We are interested in developing integrative approaches to model and predict these downstream effects of mutations in cancer.